How to Evaluate AI-Generated Data
What “good” looks like in the age of generative biology
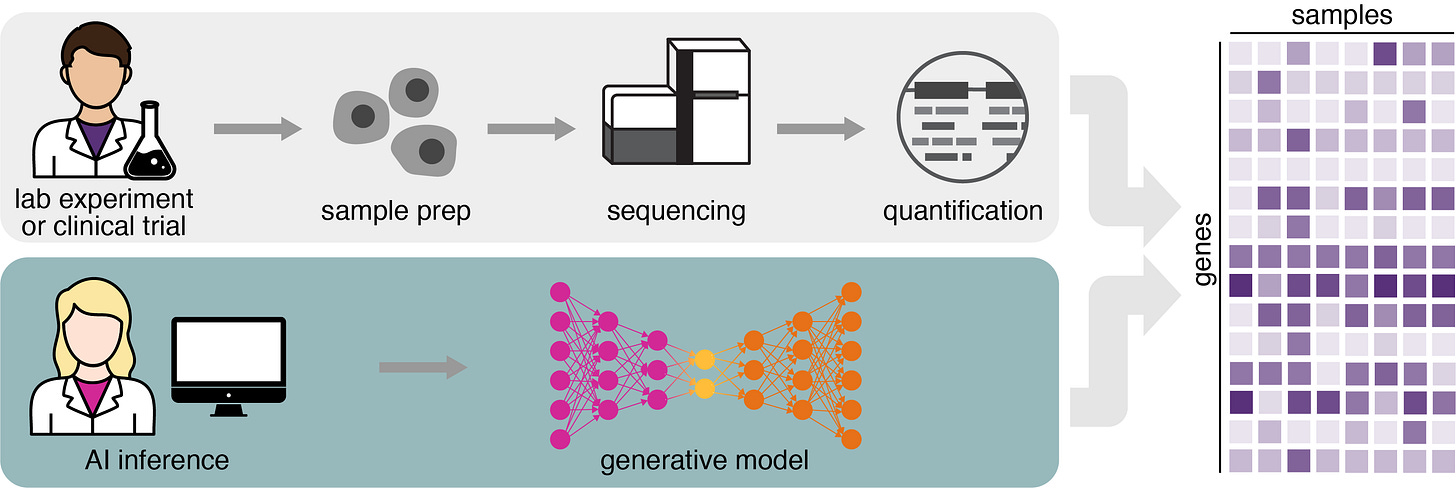
Our first model, GEM-1, simulates experiments to generate realistic gene expression data, which opens the door to entirely new kinds of discovery, significantly lower experimental costs, and lower trial failure rates. However, as AI-generated data becomes indistinguishable from lab-produced data, we face a new critical challenge:
How do we rigorously evaluate AI-generated biological data?
Traditional machine learning metrics, such as Mean Squared Error (MSE) are insufficient for two main reasons:
Variability of Ground Truth: Biological ground truth itself varies across labs, platforms, and sample-handling conditions. There’s no single correct answer to benchmark against.
Dimensionality and Structure: Gene expression data is high-dimensional (~44K genes are quantified in our bulk data models) and highly structured, with strong biological covariance patterns that inflate naive similarity metrics. Standard measures often inflate similarity because transcriptomes share an underlying structure, making even unrelated samples appear similar.
To address this, we use five core principles to evaluate the integrity and validity of AI-generated data.
Core principles for evaluating AI-generated data
I. Evaluate against laboratory reproducibility, not perfection
Just as two laboratory experiments do not yield exactly identical results, AI-generated data cannot match a laboratory-measured dataset perfectly, because every measurement contains noise arising from technical and biological variability. Complex models can learn optimized and denoised representations of the data that can efficiently capture true biological signals.
So instead of comparing AI outputs to a theoretical ideal, we should be comparing them to actual reproducibility benchmarks.
Replicates: Samples generated in the same study, under identical conditions. These represent the best possible agreement you can expect between experiments.
Pseudoreplicates: Samples from different studies with identical metadata (i.e., same cell line, same perturbation, etc). These represent reproducibility across labs, capturing variation in conditions or protocol.
Unrelated Samples: A negative control to show whether the model is capturing meaningful structure or just similarities in background. Samples match only on broad species level constraints.
If the AI-Generated data is able to reach pseudoreplicate level reproducibility, it means that it is performing as well as biology itself does across labs. Replicate-level performance — especially when we condition on reference samples — means that the AI-generated data is able to achieve the best possible fidelity that any experiment can.
II. Use metrics that match the biology
Gene expression models like GEM-1 must be evaluated not only by global similarity metrics, but also by metrics that reflect the structure and variability of the transcriptome.
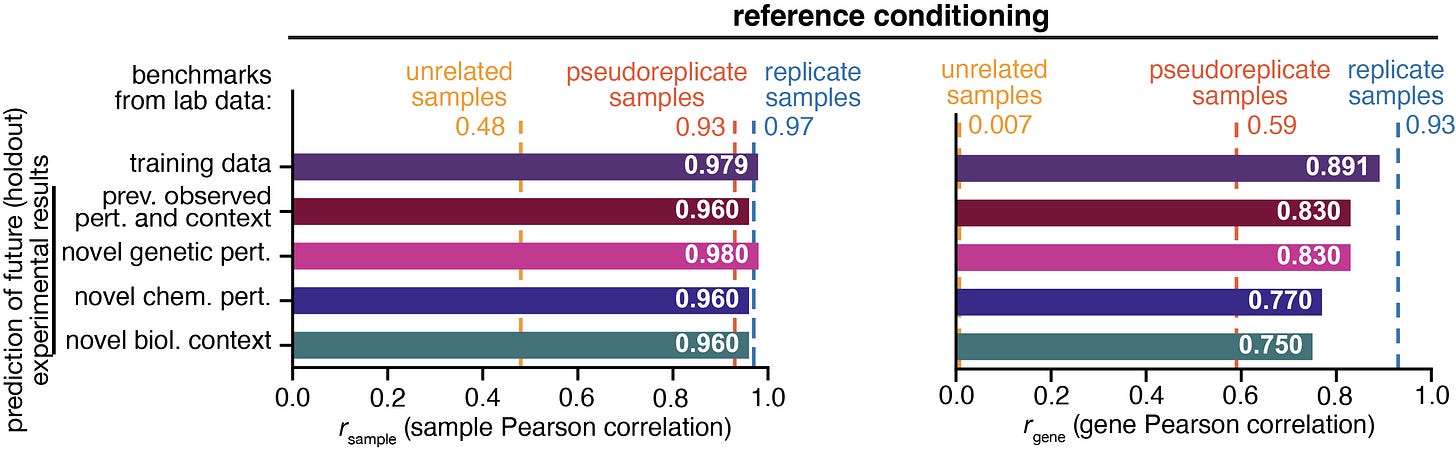
Sample Pearson Correlation: Measures how similar two samples are. But, because whole transcriptome profiles are structured, even unrelated samples show high correlations. This compresses the dynamic range of the metric.
For the above graph, the sample Pearson equivalents are:
for unrelated samples: 0.48
for pseudoreplicate samples: 0.93
for replicate samples: 0.97
Gene Pearson Correlation: Evaluates the model’s ability to rank each gene across samples. This penalizes models that merely capture baseline gene expression structure and rewards models that capture biologically meaningful variation.
The gene Pearson metric behaves more like a real test of biological inference. It asks whether the model understands how genes move relative to each other across conditions. As you can see in the figure above, the gene Pearson metric is better calibrated for model evaluation. The unrelated samples benchmark is near zero, and the replicate samples benchmark is near one.
III. Evaluate differential expression, not just expression levels
Focusing on how expression changes and not just absolute values reveals where the model captures mechanistically meaningful biology. Looking at the rank agreement of DE genes and using pathway enrichment enables us to ask questions like:
Is a drug’s AI predicted signature enrichment for the same pathways observed in wet lab data?
Are the correct stress, immune, metabolic, or signaling pathways activated?
IV. Filling in metadata
One less obvious but powerful benefit of GEM-1 is that it was trained jointly with a metadata classifier. During training, GEM-1 is not only learning how to reconstruct gene expression but also learning how to answer questions like:
Which cell line is this expression profile likely to have come from?
Does this look like a primary tissue or a cell line?
What sex, tissue, disease state, or age does this sample resemble?
This multitask design allows GEM-1 to sharpen and disambiguate the latent space, reducing the impact of technical variation on the latent space by forcing latent representations to separate metadata cleanly.
It also allows us to quantify how well the model understands any given sample. Because the classifier outputs predicted metadata with confidence scores, we can directly evaluate how coherently the model interprets an expression profile.
In addition to providing a measure of confidence, this can support use cases like QC, sample provenance, and metadata correction.
V. Validate against large real-world datasets
When generating synthetic clinical cohorts (e.g., healthy tissues, autoimmune disease cases, or cancer samples), we evaluate:
Whether tissues cluster correctly
Whether disease signatures replicate known biology
Whether individual marker genes behave as expected
Whether known subtype structures (e.g., cancer CMS subtypes) emerge naturally
Whether expression distributions match data from clinical cohorts like GTEx or TCGA
This helps establish whether the model captures patient-level variability and biological programs, not just “average” gene expression.
The Takeaway
AI-generated biological data should be evaluated using the same standards that we use to evaluate laboratory and clinical experiments. Because we know biology to be noisy, heterogeneous and context dependent, the goal of GEM-1 is to match the fidelity and variability of experimental systems and reproduce important biological signals.
When we evaluated GEM-1 with replicate-level benchmarks, pathway-level validation, metadata reconstruction, and assessments of the quality of its generated cohorts, GEM-1 proved its ability to serve as a genuine experimental engine.
And that’s where things get exciting – when AI-generated experiments behave like laboratory or clinical studies, we unlock a fundamentally new way to do science!
Learn more
Read the preprint: https://doi.org/10.1101/2025.09.08.674753
Try out GEM-1: https://app.synthesize.bio/
Take a quiz to test whether you can tell the difference between laboratory- and AI-generated data: https://synthesizebio.substack.com/p/can-you-spot-the-difference-between